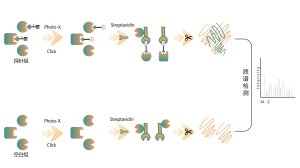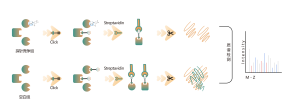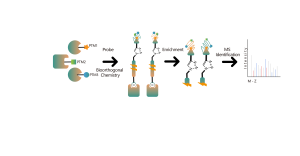Products
Quantitative Analysis of Occupancy and Selectivity of Covalent Small Molecule Drug Targets
Covalent drugs primarily work by forming covalent bonds with specific amino acid residues on target proteins, such as cysteine, lysine, and serine. Aspirin is one of the earliest known covalent drug molecules. Additionally, many natural products exhibit covalent properties, such as oridonin, which has anti-inflammatory bioactivity. In recent years, covalent targeted drugs have gained increasing attention from pharmaceutical companies. To date, at least six covalent drugs targeting kinases have been approved by the FDA, including ibrutinib, which targets the BTK kinase.
In the discovery of covalent drug targets in cells or tissues, an effective strategy is the chemical modification of active small molecules by introducing reporter groups (such as biotin or bioorthogonal groups). This modification, while maintaining the molecule’s original activity, allows for the direct capture of interacting protein targets in live cells or tissues. However, many active molecules are challenging to modify chemically, or the products formed after reacting with amino acid residues are unstable, making them unsuitable for mass spectrometry detection. To overcome these challenges, ChomiX offers a solution: modification site identification using competitive labeling with amino acid-specific probes.
Technical Platform
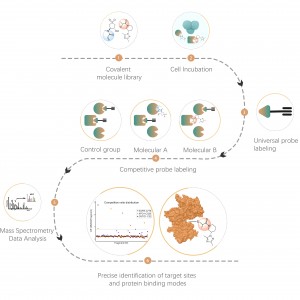
Our technology platform is centered around a universal aminoacids-specific chemical probe. When an active small molecule reacts with an amino acid residue and occupies the binding site, this universal chemical probe generates a significant signal difference for that binding site compared to blank control samples. By detecting these differences in the labeling signals, we can accurately obtain information about the target proteins and amino acid residues of the active molecule, including both expected (on-target) and potential off-target proteins. This technology platform provides robust support for target discovery of covalent drugs and research into drug mechanisms of action.

Our Advantages
1. Technical Excellence: Experienced team, top-tier journal publications, and authoritative industry services.
2. Core Patent Technology: Exclusive patents and advanced hardware for early drug development support.
3. One-stop Service: Covering probe design, synthesis, target discovery, bioinformatics analysis, and timely progress feedback for customer satisfaction.
4. Rigorous Quality Management: ISO9001 certification ensures trustworthy and authentic reports.
Our Service
| Project | Quantitative Analysis of Occupancy and Selectivity of Covalent Small Molecule Drug Targets |
| Sample | Pure protein, cell lysate, live cells, diseased tissue, blood, bacteria, plant tissue |
| Hardware Platform | Non-contact ultrasonic cell pulverizer,ChemiDoc MP Imaging System,Orbitrap Fusion Lumos Tribrid/Orbitrap Exploris 480/Q Exactive HF-X/timsTOF Pro 2 mass spectrometer |
| Project Duration | 2-4 weeks |
| Deliverables | Project Report (including experimental procedures, data analysis charts, bioinformatics analysis results) |
| Price | Click to consult |
Case Study
AMG510, developed by Amgen, is the world's first targeted drug for KRAS-G12C mutant tumors. This project aims to verify its target specificity and selectivity in corresponding mutant cells. Using Chomix's DIA-ABPP platform, we comprehensively screened the covalent targets of AMG510 in cells down to the amino acid residue level.
Experimental data show that in four repeated experiments on NCI-H358 cells, a total of 16,992 cysteine residues from 5,768 proteins were systematically analyzed. Under 1μM AMG510 treatment, the KRAS_C12 site showed significant changes, while KRAS_C80 remained unaffected, providing strong evidence for the high specificity of AMG510 towards the KRAS-G12C mutant site (marked with an asterisk indicating the targeted cysteine residue site).