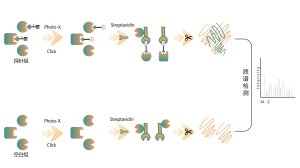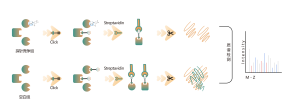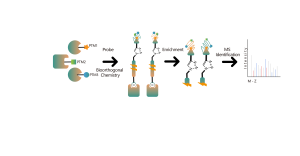Products
Identification of the binding site for non-covalent small molecule drugs in living cells
At the cellular level, directly studying the mode of action between drugs and target proteins can help avoid changes in protein structure during purification and false-positive results that may arise from artificial buffer systems and high drug-protein concentrations.
Technical Platform
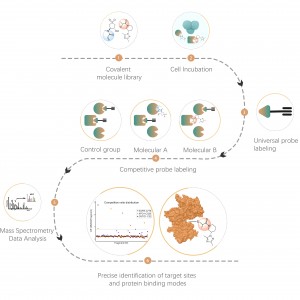
ChomiX provides chemical proteomics services utilizing biologically active photoreactive chemical probes (with activity similar to drug molecules). These probes are incubated with disease-relevant cells or tissues at physiologically relevant concentrations of drugs, followed by in situ rapid photo-crosslinking technology to convert non-covalent interactions between drug molecule probes and protein targets into covalent interactions. Subsequently, through steps such as target protein enrichment, enzymatic digestion to release unmodified peptide segments, and selective enrichment of drug molecule probe-modified peptide segments, combined with high-resolution mass spectrometry of biomolecules, peptide sequence information can be rapidly determined. Finally, molecular docking tools are employed to quickly obtain binding models of drugs with target proteins, providing robust support for subsequent medicinal chemistry research.

Our Advantages
1. Technical Excellence: Experienced team, top-tier journal publications, and authoritative industry services.
2. Core Patent Technology: Exclusive patents and advanced hardware for early drug development support.
3. One-stop Service: Covering probe design, synthesis, target discovery, bioinformatics analysis, and timely progress feedback for customer satisfaction.
4. Rigorous Quality Management: ISO9001 certification ensures trustworthy and authentic reports.
Our Service
| Project | Identification of Binding Pockets for Non-covalent Small Molecule Drugs |
| Sample | Pure protein, cell lysate, live cells, diseased tissue, blood, bacteria, plant tissue |
| Hardware Platform | Non-contact ultrasonic cell pulverizer,ChemiDoc MP Imaging System,Orbitrap Fusion Lumos Tribrid/Orbitrap Exploris 480/Q Exactive HF-X/timsTOF Pro 2 mass spectrometer |
| Project Duration | 2-4 weeks |
| Deliverables | Project Report (including experimental procedures, data analysis charts, bioinformatics analysis results) |
| Price | Click to consult |
Case Study
The target of candidate drug molecule B is a multi-transmembrane protein with multiple drug binding pockets. Despite multiple attempts using structural biology methods such as X-ray and cryo-EM, a binding model between the drug molecule and the target protein could not be obtained.
Based on the structure and activity of candidate drug molecule B, our company designed and synthesized a photoreactive chemical probe, Probe B, comprising photoreactive and bioorthogonal groups. Utilizing the chemical proteomics target discovery platform mentioned above, target proteins were first validated in cell lines relevant to the activity of candidate drug molecule B. Subsequently, leveraging mass spectrometry-based non-covalent drug binding pocket identification technology, the spatially adjacent peptide segments to the drug binding pockets were determined, and a drug-protein binding model was provided.

Immunoblotting assays revealed that candidate drug molecule B effectively competes with probe labeling signals in cells, indicating direct binding between the candidate drug molecule and the target protein.

As chemical probes can only crosslink with peptide segments in close spatial proximity, the sequence CLPFIIGCNPTILHVHELYIR was identified through tandem high-resolution mass spectrometry to determine the crosslinked peptide segments.